Modeling of snRNP Motion in the Nucleoplasm
Small nuclear ribonucleoprotein particles (snRNPs) are essential supramolecular complexes involved in pre-mRNA splicing, the process of post-transcriptional RNA modifications. The particles undergo complex assembly steps inside the cell nucleus in a highly dynamic compartment called the Cajal body.
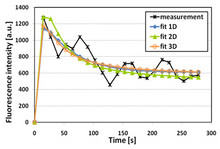
We have previously shown that the free diffusion model does not fully describe the snRNP motion when tested for 1D fundamental solution of the diffusion equation. Instead, here we use numerical solutions of the diffusion equation to fit the experimental data obtained by fluorescence microscopy of a single cell in order to test the influence of geometry, dimensionality and boundary conditions. Finally, optimal characteristics of the model were determined.
Download
- Blazikova_pres.pdf - 0.52MB
- Blazikova.pdf - 0.55MB



